Epigenetics
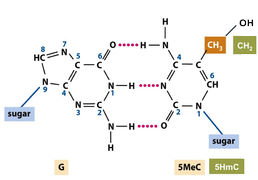
Epigenetics referes to the study of biological processes that regulate gene transcription within the cell that are heritable mitoticaly and/or meioticaly but that do not involve changes in the DNA sequence. This include regulation of DNA packaging within the nucleus through chromatin remodelling involving histone modifications and chemical changes to the DNA itself such as cytosine methylation.
Within the Faculty of Medicine there are several groups generating large data sets assessing DNA methylation genome-wide utilising both array based platforms and direct sequencing of bisulphite converted DNA. Analysis of DNA methylation variation between individuals in relation to disease state adds an order of magnitude in complexity to genome wide association analysis of DNA sequence variation as methylation values are quantitative rather than binary and vary over time, between tissues and with environmental exposure.
For queries about this topic, contact John Holloway.
View the calendar of events relating to this topic.
Projects

Centre for Doctoral Training in Next Generation Computational Modelling
Hans Fangohr, Ian Hawke, Peter Horak (Investigators), Susanne Ufermann Fangohr, Thorsten Wittemeier, Kieran Selvon, Alvaro Perez-Diaz, David Lusher, Ashley Setter, Emanuele Zappia, Hossam Ragheb, Ryan Pepper, Stephen Gow, Jan Kamenik, Paul Chambers, Robert Entwistle, Rory Brown, Joshua Greenhalgh, James Harrison, Jonathon Waters, Ioannis Begleris, Craig Rafter
The £10million Centre for Doctoral Training was launched in November 2013 and is jointly funded by EPSRC, the University of Southampton, and its partners.
The NGCM brings together world-class simulation modelling research activities from across the University of Southampton and hosts a 4-year doctoral training programme that is the first of its kind in the UK.
Identifying factors required for DNA methylation using the imprinting control protein ZFP57
Deborah Mackay (Investigator)
Mutation of ZFP57 in humans is associated with widespread loss of DNA methylation at imprinted genes, and clinical features including congenital anomalies and developmental delay (Mackay et al, 08). This indicates that ZFP57 is required for DNA methylation of imprinted genes necessary for normal development.
We propose to identify the DNA sequences targeted by ZFP57, and its protein cofactors. This work will give insight into the biology of imprinting, indicate mechanisms of disease, and identify novel imprinted genes.
.small.png)
Imprinting Disorders Finding Out Why
Karen Temple (Investigator)
We are conducting a research project to determine the cause and clinical impact of widespread imprinting aberrations in human development. We are recruiting patients with possible or definite imprinting disorders (due to methylation loss or gain at an imprinted loci)
including Silver Russell syndrome, Transient Neonatal diabetes, Beckwith Wiedemann syndrome, Angelman syndrome Prader Willi syndrome, UPD 14 syndromes and Pseudohypoparathyroidism.

Mathematical tools for analysis of genome function, linkage disequilibrium structure and disease gene prediction
Mahesan Niranjan, Andrew Collins, Reuben Pengelly (Investigators)
This iPhD project uses a Gaussian Bayesian Networks framework through Machine learning methods to predict which genes are involved in the development of different diseases.

Mathematical tools for analysis of genome function, linkage disequilibrium structure and disease gene prediction
Mahesan Niranjan, Andrew Collins, Reuben Pengelly (Investigators)
This PhD project uses a Monte Carlo molecular simulation processes approach to predict which genes are involved in the development of different diseases.

Transgenerational inheritance of allergy in a multi generational cohort
John Holloway (Investigator)
The aim of this project is to determine the vertical transmission of DNA methylation by identification of CpG sites by microarray analysis of 450,000 CpG sites in 252 women of the IoW cohort that are associated with allergic sensitization and testing whether the identified methylation patterns are vertically transmitted to their offspring and whether modifiable environmental conditions during gestation affect DNA methylation.
People
 Andrew Collins
Andrew CollinsProfessor, Medicine (FM)
 Hans Fangohr
Hans FangohrProfessor, Engineering Sciences (FEE)
 John Holloway
John HollowayProfessor, Medicine (FM)
 Mahesan Niranjan
Mahesan NiranjanProfessor, Electronics and Computer Science (FPAS)
 Karen Temple
Karen TempleProfessor, Medicine (FM)
 Peter Horak
Peter HorakReader, Optoelectronics Research Centre
 Deborah Mackay
Deborah MackayReader, Medicine (FM)
 Reuben Pengelly
Reuben PengellySenior Lecturer, Medicine (FM)
 Christopher Bell
Christopher BellLecturer, Biological Sciences (FNES)
 Ian Hawke
Ian HawkeLecturer, Mathematics (FSHS)
 Gaia Andreoletti
Gaia AndreolettiPostgraduate Research Student, Medicine (FM)
 Alice Ball
Alice BallPostgraduate Research Student, Electronics and Computer Science (FPAS)
 Ioannis Begleris
Ioannis BeglerisPostgraduate Research Student, Engineering Sciences (FEE)
 Rory Brown
Rory BrownPostgraduate Research Student, Civil Engineering & the Environment (FEE)
 Paul Chambers
Paul ChambersPostgraduate Research Student, Engineering Sciences (FEE)
 Caroline Duignan
Caroline DuignanPostgraduate Research Student, Biological Sciences (FNES)
 Robert Entwistle
Robert EntwistlePostgraduate Research Student, Engineering Sciences (FEE)
 Stephen Gow
Stephen GowPostgraduate Research Student, Engineering Sciences (FEE)
 Joshua Greenhalgh
Joshua GreenhalghPostgraduate Research Student, Engineering Sciences (FEE)
 James Harrison
James HarrisonPostgraduate Research Student, Engineering Sciences (FEE)
 William Hurndall
William HurndallPostgraduate Research Student, Electronics and Computer Science (FPAS)
 Konstantinos Kouvaris
Konstantinos KouvarisPostgraduate Research Student, Electronics and Computer Science (FPAS)
 David Lusher
David LusherPostgraduate Research Student, Engineering Sciences (FEE)
 Gregory Parkes
Gregory ParkesPostgraduate Research Student, Electronics and Computer Science (FPAS)
 Alvaro Perez-Diaz
Alvaro Perez-DiazPostgraduate Research Student, Engineering Sciences (FEE)
 Daniel Power
Daniel PowerPostgraduate Research Student, Electronics and Computer Science (FPAS)
 Craig Rafter
Craig RafterPostgraduate Research Student, Engineering Sciences (FEE)
 Hossam Ragheb
Hossam RaghebPostgraduate Research Student, Engineering Sciences (FEE)
 Sonya Ridden
Sonya RiddenPostgraduate Research Student, Mathematics (FSHS)
 Kieran Selvon
Kieran SelvonPostgraduate Research Student, Engineering Sciences (FEE)
 Ashley Setter
Ashley SetterPostgraduate Research Student, Engineering Sciences (FEE)
 Jonathon Waters
Jonathon WatersPostgraduate Research Student, Engineering Sciences (FEE)
 Thorsten Wittemeier
Thorsten WittemeierPostgraduate Research Student, Engineering Sciences (FEE)
 Emanuele Zappia
Emanuele ZappiaPostgraduate Research Student, Engineering Sciences (FEE)
 Matthew Higgins
Matthew HigginsUndergraduate Research Student, Biological Sciences (FNES)
 Shaun Maguire
Shaun MaguireUndergraduate Research Student, Biological Sciences (FNES)
 Susanne Ufermann Fangohr
Susanne Ufermann FangohrAdministrative Staff, Civil Engineering & the Environment (FEE)
 Jan Kamenik
Jan KamenikAlumnus, University of Southampton
 Brian Bonney
Brian BonneyNone, None
 Roshan Sood
Roshan SoodNone, None