Antimicrobial Peptide and E. coli Membrane Interactions
- Research Team
- Thomas Piggot, Nils Berglund
- Investigators
- Syma Khalid
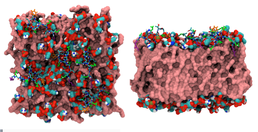
With an alarming number of bacterial strains showing resistance to modern antibiotic treatments it is essential that we advance in the field of novel antibiotic research. Through investigating the mechanism by which naturally occurring antibacterial proteins disrupt bacterial membranes we can discover new ways of fighting bacteria I work with two antimicrobials, Melittin and Polymyxin B1 (PMB). Although it is known that these AMPs permeate the bacterial cell membrane, the mechanism through which it does so is still in debate. Using the molecular dynamics simulation software GROMACS, I can simulate the interaction between the AMPs and cell membrane and study the interactions on an atomic level. Through our research we have been able to determine that the addition of PMB to the bacterial inner membrane (IM) causes changes in its physical properties; we suspect that these changes are key to the disruption of the membrane. We have also looked at the stability of melittin pores in different cell membranes, allowing us to determine the mechanisms by which these pores function and kill different cells. Thanks to Iridis we can simulate on a ?s timescale allowing us to see the insertion of the AMPs in the bacterial membrane and gain a deeper understanding of these biological systems.
Categories
Life sciences simulation: Bioinformatics, Biomolecular simulations
Simulation software: Gromacs
Visualisation and data handling software: VMD
Computational platforms: Iridis, Mac OS X
Transdisciplinary tags: Complex Systems, Scientific Computing