Seminar 13th May 2011 3:30 p.m. University of Southampton, Building 58 (Social Sciences) Room 1007
Molecular modeling - now fun and colorful!
Dr Marc Baaden
Laboratoire de Biochimie Théorique CNRS - UPR9080 Institut de Biologie Physico-Chimique
- Web page
- http://www.baaden.ibpc.fr/
- Categories
- AMBER, Biomechanics, C, C++, CASTEP, Complex Systems, Computer Science, COMSOL, Density functional Theory, e-Research, Education, FFT, Fortran, GPU, GPU-libs, HECToR, HPC, Materials, Mathematica, Metals, Molecular Dynamics, Molecular Mechanics, Monte Carlo, MPI, Multi-physics, Multi-scale, Onetep, PETSc, Photonics, Quantum Chemistry, Scientific Computing, Semiconductors, Sensors, Software Engineering, Tribology, Xmgrace
- Submitter
- Chris-Kriton Skylaris
Complex Systems Simulation Seminar Series (CS^4)
from the Institute for Complex Systems Simulation, the Complexity in Real-World Contexts USRG, the Computational Modelling Group and the Computational Systems Chemistry section (School of Chemistry).
Abstract
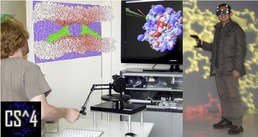
Studying macromolecular assemblies interactively is becoming an increasingly appealing approach to molecular modeling [1]. Interactive simulations are precious in exploring and generating hypotheses. They introduce a human factor and benefit from the user's experience and insight. By using a 3D haptic device, molecules can be manipulated with great precision and interactions can be felt by the user in real time via tactile feedback.
With a focus on interactive approaches to study macromolecular structure, flexibility and interactions, I will describe several investigations of enzymatic systems and membrane proteins. In this context, visualisation can be another limiting factor, and I will illustrate the possibilities offered by novel GPU implementations of molecular structure depiction with methods such as shader-based ray-casting [2].
[1] Delalande & al, Complex Molecular Assemblies at hand via Interactive Simulations, J. Comput. Chem. 30, 2009, 2375-2387 [2] Chavent & al, GPU-powered tools boost molecular visualization, Brief. Bioinf., doi:10.1093/bib/bbq089 (online)
Refreshments
Available from 3:30pm, lecture starts at 4pm.
Contact
Multidisciplinary Research Co-ordinator
University Strategic Research Groups
Research and Innovation Services
02380 593244