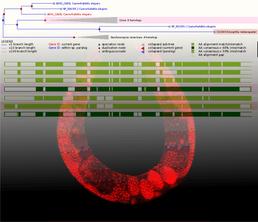
Bioinformatic identification and physiological analysis of ethanol-related genes in C. elegans
- Homepage
- http://www.southampton.ac.uk/song/
- Started
- 1st October 2008
- Research Team
- Ben Ient
- Investigators
- Richard Edwards, Vincent O'Connor, Lindy Holden-Dye
There is an urgent need to better understand the biological mechanism that underpins addiction. When factoring health and societal burdens, alcohol is recognised amongst the most harmful addictive drugs. Despite its simple structure, alcohol is known to impact both acutely and chronically on several molecules (e.g. ion channels, transmitter receptors, signalling enzymes) to modify the circuits that control complex behaviour. In humans, complex interactions between these molecular, cellular and systems level targets underlie the euphoric, depressive, and anaesthetic effects of alcohol and the more complex outcomes of tolerance, withdrawal and drug seeking.
This multidisciplinary project investigates the broad molecular, cellular and systems level impacts of acute and chronic ethanol in the nematode, Caenorhabditis elegans, as a model; many of the mammalian molecular and cellular targets implicated in the action of alcohol are conserved in this organism. The work involves building a database in which key facets of ethanol and its targets are collated and from which a comparative genomics bioinformatic approach is being used to predict key molecular targets of ethanol action for experimental investigation in C. elegans.
Categories
Life sciences simulation: Bioinformatics, Biomedical, Biomolecular Organisation, Neuroscience, Systems biology
Visualisation and data handling methods: Database
Visualisation and data handling software: MS Office Access